If you are like me and have been using R for a long time but would like to explore and add some python capabilities to your workflows, reticulate + R-Studio is a great way to achieve just that.
Here’s a quick excerpt from reticulate’s website:
The reticulate package provides a comprehensive set of tools for interoperability between Python and R.
Essentially, reticulate allows R to talk to Python (via a live python session running in the background) and works seamlessly within RStudio. It also provides functionality to manage multiple python installations. In this post, we’ll explore how to set up a python environment and configure the same to work with RStudio in windows. Let’s dive in!
Installing Python
The first step, ofcourse, is to install the reticulate package.
install.packages("reticulate")
Next, we will install python via reticulate. For windows, if installing python via reticulate, it’s better to install miniconda since reticulate only uses conda to install and manage python libraries. If you’d rather use a bare bones installation of python, it would be a lot easier to directly install python using the windows binary, go through the installation process, use pip from either the command prompt or Power Shell to install the libraries you need and then point reticulate to that installation of python using use_python(). See here for more details.
This post will use Miniconda. By default, a virtual environment called r-reticulate will also be created as part of the installation process.
library(reticulate)
install_miniconda(path = "e:/miniconda", update = T)
Pointing to the right python installation
Now that Miniconda is installed, we need to point reticulate to it. First, let’s check if the r-reticulate virtual environment is available.
conda_list(conda = "e:/miniconda/_conda.exe")
## name python
## 1 r-reticulate E:\\miniconda\\envs\\r-reticulate\\python.exe
If available, then point to it using use_condaenv(). This binds the particular installation of Python to the current R session.
Note that if you restart R, this will need to be set again. To set it permanently set the RETICULATE_PYTHON environment variable using Sys.setenv().
use_condaenv(condaenv = "r-reticulate", conda = "e:/miniconda/_conda.exe")
If the default virtual environment is not available, or you would like to create a new one, then use conda_create()
conda_create(envname = "myenv", conda = "e:/miniconda/_conda.exe")
Installing libraries
Now that reticulate nows where to find python, we can install some python libraries to work with.
py_install(packages = c("pandas", "scikit-learn"))
Using Python within R-Studio
If all of the above steps worked without any errors, you should be able to do something like this in a new R session (console or R-script).
pd <- import("pandas")
pd$array(c(1, 2, 3))
## <PandasArray>
## [1.0, 2.0, 3.0]
## Length: 3, dtype: float64
Yay! Python and R are now talking to each other 👏
Now, that everything is setup, there are multiple ways to use python along with R inside RStudio:
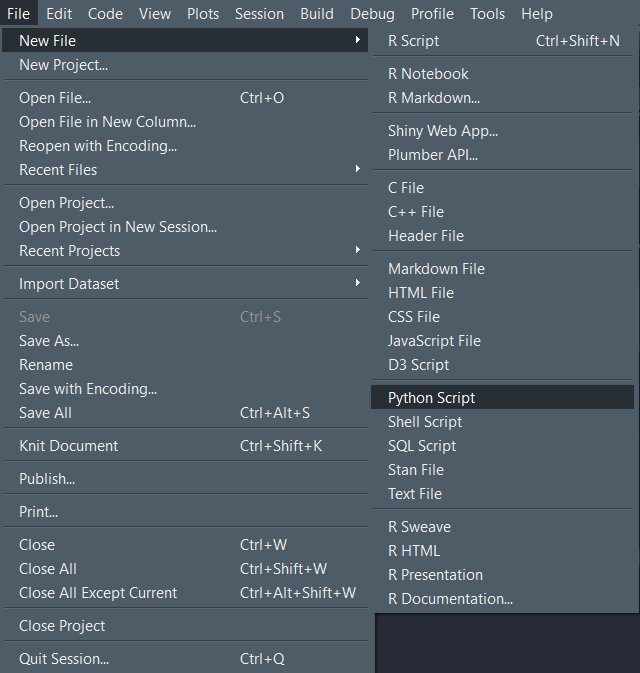
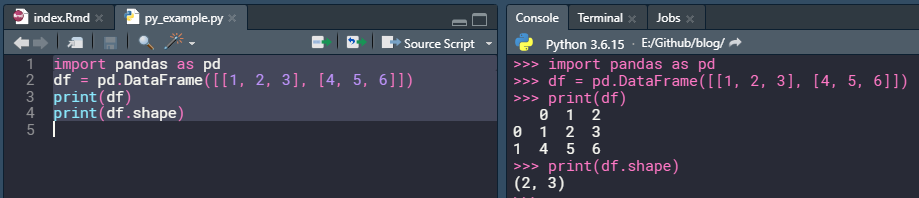
One can use R-Studio as an IDE for python. Simply open up a new python script from File -> New File -> Python Script and start to write some python code.
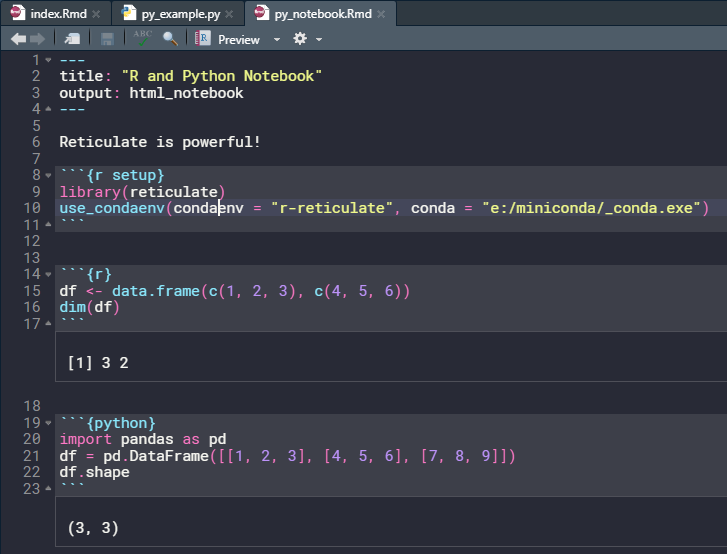
Another way to use python inside RStudio is via R-Notebooks. Simply launch a new R-Notebook and start to write python code inside a python code chunk. This way, one can use Python and R within the same notebook. 😏
Possibly the most exciting way of using Python with R is to import Python functions into R. This is a great way to add python functionality to an existing R environment.
- We’ll need a python script to house python code including functions that need to be imported into R
- We can then import the above functions via
source_python()
As an example, say we have a python script called py_example.py with the following code which allows us to fit a linear regression model using scikit-learn.
from sklearn import linear_model
linreg_python = linear_model.LinearRegression()
We can import this function into R by simply running:
source_python("py_example.py")
And now we can use this function inside R as we would use any other function (note that syntax is different however).
# Fit model
linreg_python$fit(X = mtcars[,-1], y = mtcars$mpg)
## LinearRegression(copy_X=True, fit_intercept=True, n_jobs=None, normalize=False)
# Show coefficients
data.frame(var = c("Intercept", names(mtcars)[-1]),
python_coef = c(linreg_python$intercept_, linreg_python$coef_))
## var python_coef
## 1 Intercept 12.30337416
## 2 cyl -0.11144048
## 3 disp 0.01333524
## 4 hp -0.02148212
## 5 drat 0.78711097
## 6 wt -3.71530393
## 7 qsec 0.82104075
## 8 vs 0.31776281
## 9 am 2.52022689
## 10 gear 0.65541302
## 11 carb -0.19941925
Just for fun, let’s also compare the output from good old lm()
# Fit model and show coefficients from R
fit <- lm(mpg ~ ., data = mtcars)
data.frame(R_coef = coef(fit))
## R_coef
## (Intercept) 12.30337416
## cyl -0.11144048
## disp 0.01333524
## hp -0.02148212
## drat 0.78711097
## wt -3.71530393
## qsec 0.82104075
## vs 0.31776281
## am 2.52022689
## gear 0.65541302
## carb -0.19941925
Using the above set-up, one could combine python’s extensive ML capabilities and R’s intuitive data munging and excellent data visualisation capabilities into one single powerful workflow. I wonder if Py-ThoR would be an appropriate name for such a workflow? 👊
Thoughts? Comments? Helpful? Not helpful? Like to see anything else added in here? Let me know!